An Overview of Identification Methods on Human Brain Effective Connectivity Networks Based on Functional Magnetic Resonance Imaging
-
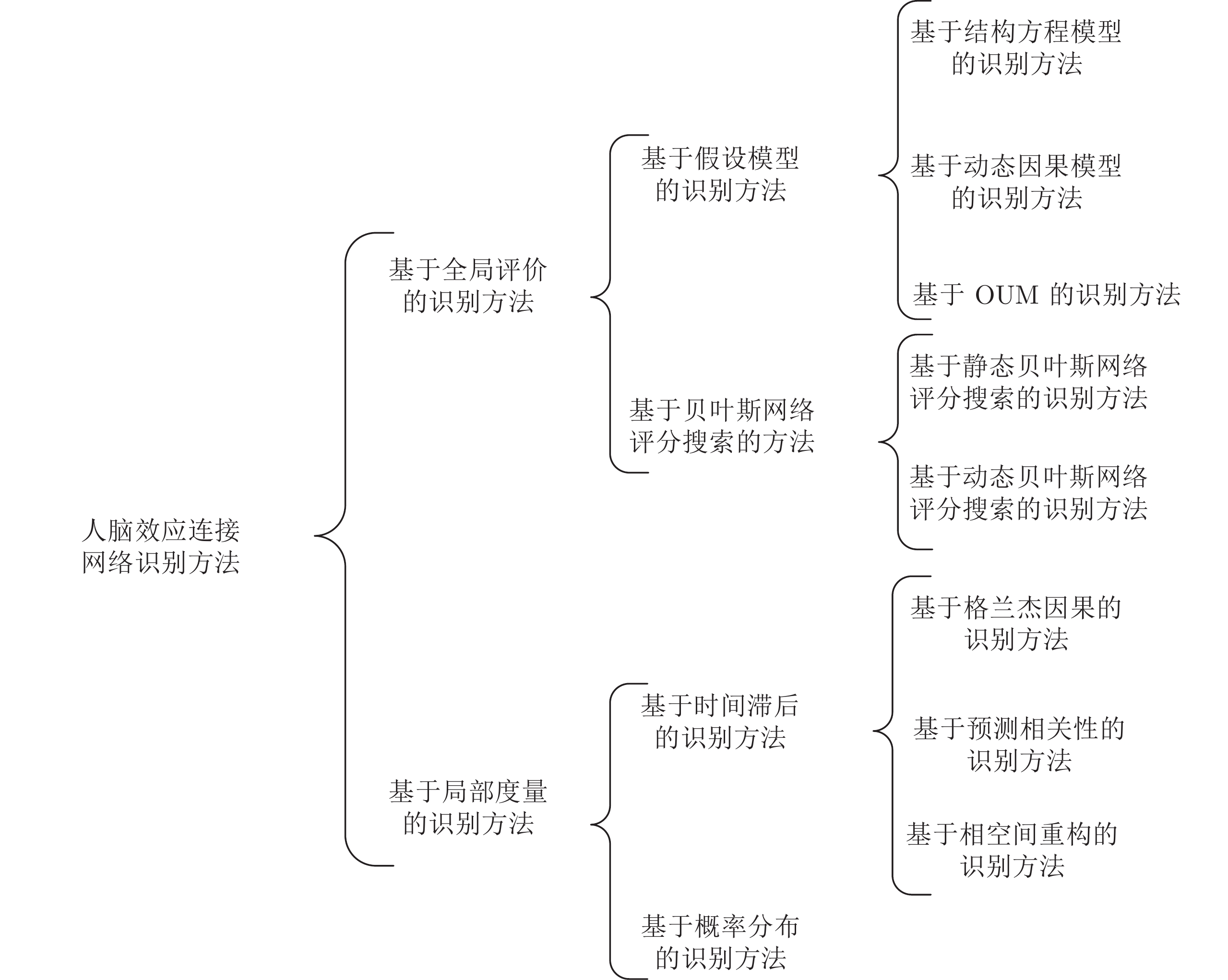
摘要: 人脑效应连接网络刻画了脑区间神经活动的因果效应. 对不同人群的脑效应连接网络进行研究不仅能为神经精神疾病病理机制的理解提供新视角, 而且能为疾病的早期诊断和治疗评价提供新的脑网络影像学标记, 具有十分重要的理论意义和应用价值. 利用计算方法从功能磁共振成像(Functional magnetic resonance imaging, fMRI)数据中识别脑效应连接网络是目前人脑连接组学中一项重要的研究课题. 本文首先概括了从fMRI数据中进行脑效应连接网络识别的主要流程, 说明了其中的主要步骤和方法; 然后, 给出了一种脑效应连接网络识别方法的分类体系, 并对其中一些代表性的识别算法进行了阐述; 最后, 通过对该领域挑战性问题的分析, 预测了脑效应连接网络识别未来的研究方向, 以期对相关研究提供一定的参考.Abstract: The brain effective connectivity networks characterize the causal interactions of neural activity between brain regions. Researches on brain effective connectivity networks of different populations can not only provide a new perspective for understanding the pathological mechanism of neuropsychiatric diseases, but also provide novel brain network imaging markers for the early diagnosis and evaluation for treatment of diseases, thus have very important theoretical and practical value. Using computational approaches to identify brain effective connectivity networks from functional magnetic resonance imaging (fMRI) data is currently an important subject in the human brain connectome. This paper firstly summarizes a workflow of identifying brain effective connectivity networks from fMRI data and illustrates its main processes and methods. Next, a comprehensive category system of identifying brain effective connectivity networks is presented, and several typical identifying algorithms in each category are described. Finally, by analyzing challenging problems in this area, we predict the further research directions in identifying brain effective connectivity networks and hope to present some references for related researches.
-
表 1 人脑效应连接网络的部分典型识别方法对比
Table 1 The comparisons of several typical identification methods on human brain effective connectivity networks
方法 年份 类别 个体水平/组水平 探索型/验证型 主要优缺点及适用场景 uSEM[55] 2007 基于结构方程模型 个体水平 验证型 适用于组块设计实验, 能够识别同期和滞后的因果效应, 但不能用于事件相关的实验设计 euSEM[56] 2011 基于结构方程模型 个体水平、组水平 验证型、探索型 适用于事件相关设计实验, 不仅识别了同期和滞后的因果效应, 而且描述了外界刺激对脑区神经活动的直接影响和间接调节作用, 在个体水平和组水平上都获得了较好的识别效果 GIMME[58] 2012 基于结构方程模型 个体水平、组水平 验证型、探索型 从高度异质数据中准确地识别了组水平和个体水平的效应连接网络, 且具有良好的灵活性和可扩展性 cs-GIMME[59] 2019 基于结构方程模型 个体水平、组水平 验证型、探索型 与 GIMME 相比, 具有更强地识别组间差异的能力, 但不适于识别大规模脑效应连接网络 spDCM[62] 2014 基于动态因果模型 组水平 验证型 用于从静息态 fMRI 数据中识别高精度的效应连接网络, 对组间差异敏感, 不足在于仅考虑了较小规模的脑区, 且忽略了效应连接的动态性 rDCM[67] 2017 基于动态因果模型 个体水平 验证型 适用于识别较大规模的脑效应连接网络 spare rDCM[68] 2018 基于动态因果模型 个体水平、组水平 验证型 用于准确、高效地识别全脑的效应连接网络, 且对组间差异敏感 文献 [46] 2018 基于动态因果模型 个体水平、组水平 验证型 用于识别动态效应连接网络, 方法具有良好的可扩展性 文献 [71] 2016 基于 OUM 组水平 验证型 适用于识别较大规模的效应连接网络, 但滞后信息可能对因果效应的识别造成混淆 文献 [72] 2018 基于 OUM 组水平 验证型 利用零时滞的功能连接识别效应连接网络, 避免了滞后信息的不良影响, 但方法的鲁棒性较差 AIAEC[44] 2016 基于静态 BN 评分搜索 组水平 探索型 能够准确地识别脑区间的连接方向, 且在大规模脑区上也具有良好的性能, 缺陷是不能计算连接强度 ACOEC[45] 2016 基于静态 BN 评分搜索 组水平 探索型 能够识别连接方向并计算连接强度, 且具有良好的精度和鲁棒性, 但方法的时间复杂度较高 ACOMMEC[79] 2019 基于静态 BN 评分搜索 组水平 探索型 利用多源信息识别脑效应连接网络, 比仅利用 fMRI 数据获得了更高的求解质量和计算效率, 但不能模拟人脑的循环机制 GDBN[80] 2014 基于动态 BN 评分搜索 组水平 探索型 能够识别具有反馈机制的效应连接网络, 但该方法仅采用了一阶马尔科夫链 DBN 模型, 未考虑多个不同时刻的循环交互作用 HO-DBN-DP[81] 2017 基于动态 BN 评分搜索 组水平 探索型 能够高效地识别多个不同时刻的脑区间具有循环和反馈机制的因果效应连接 TB-based score[82] 2018 基于动态 BN 评分搜索 组水平 探索型 基于融合 fMRI 和 DTI 信息的评分函数学习网络结构, 在精度和鲁棒性上均获得了良好性能, 但不能有效地处理数据缺失问题 SVAR[92] 2015 基于格兰杰因果 组水平 探索型 利用因子降维策略获得了较高的计算效率, 适用于识别较大规模的效应连接网络, 不足在于因子模型对一些特殊脑区不适用 文献 [95] 2016 基于格兰杰因果 组水平 探索型 结合了时、频域格兰杰因果的优势, 获得了更高的准确性, 但方法对模型的阶数敏感 MKGC[90] 2017 基于格兰杰因果 组水平 探索型 适用于捕捉脑区间非线性的因果效应, 具有较强的识别出脑区之间连接的能力 lsGC[97] 2019 基于格兰杰因果 组水平 探索型 能高效地识别大规模脑区间的效应连接网络(在90、138 和 246 个节点的大规模脑区上验证了有效性) P-correlation[100] 2017 基于预测相关性 组水平 探索型 在数据非平稳的情况下能够准确地识别每对脑区间的因果效应, 具有灵敏度高、鲁棒性强的特点 GS[101] 2002 基于相空间重构 组水平 探索型 适用于识别每对脑区间的连接方向, 但有时三个指标的方向判断结果不一致 MCA[102] 2018 基于相空间重构 组水平 探索型 用于识别每对脑区间非线性的因果效应, 且在样本量较小时仍具有较高的准确率 文献 [105] 2019 基于概率分布 组水平 探索型 综合考虑了 BOLD 信号多方面的信息, 在每对脑区间方向的识别上具有良好的鲁棒性和准确性 CDD[106] 2019 基于概率分布 组水平 探索型 无需严格假设数据的分布, 能够灵活、准确地度量每对脑区间的非线性因果关系 -
[1] Lehrer J. Neuroscience: Making connections. Nature, 2009, 457(7229): 524−527 doi: 10.1038/457524a [2] 梁夏, 王金辉, 贺永. 人脑连接组研究: 脑结构网络和脑功能网络. 科学通报, 2010, 55(16): 1565−1583 doi: 10.1360/972009-2150Liang Xia, Wang Jin-Hui, He Yong. Human connectome: Structural and functional brain networks. Chinese Science Bulletin, 2010, 55(16): 1565−1583 doi: 10.1360/972009-2150 [3] Friston K J. Functional and effective connectivity: A review. Brain Connectivity, 2011, 1(1): 13−36 doi: 10.1089/brain.2011.0008 [4] Song M, Jiang T Z. A review of functional magnetic resonance imaging for Brainnetome. Neuroscience Bulletin, 2012, 28(4): 389−398 doi: 10.1007/s12264-012-1244-4 [5] Deshpande G, Li Z H, Santhanam P, Coles C D, Lynch M E, Hamann S, et al. Recursive cluster elimination based support vector machine for disease state prediction using resting state functional and effective brain connectivity. PLoS One, 2010, 5(12): e14277 doi: 10.1371/journal.pone.0014277 [6] Li X F, Coyle D, Maguire L, Watson D R, McGinnity T M. Gray matter concentration and effective connectivity changes in Alzheimer' s disease: A longitudinal structural MRI study. Neuroradiology, 2011, 53(10): 733−748 doi: 10.1007/s00234-010-0795-1 [7] Friston K J. Functional and effective connectivity in neuroimaging: A synthesis. Human Brain Mapping, 1994, 2(1−2): 56−78 doi: 10.1002/hbm.460020107 [8] Smith S M, Miller K L, Salimi-Khorshidi G, Webster M, Beckmann C F, Nichols T E, et al. Network modelling methods for fMRI. NeuroImage, 2011, 54(2): 875−891 doi: 10.1016/j.neuroimage.2010.08.063 [9] Farahani F V, Karwowski W, Lighthall N R. Application of graph theory for identifying connectivity patterns in human brain networks: A systematic review. Frontiers in Neuroscience, 2019, 13: 585 doi: 10.3389/fnins.2019.00585 [10] 左西年, 张喆, 贺永, 臧玉峰. 人脑功能连接组: 方法学、发展轨线和行为关联. 科学通报, 2012, 57(35): 3399−3413 doi: 10.1360/972012-702Zuo Xi-Nian, Zhang Zhe, He Yong, Zang Yu-Feng. The human functional connectome: Its methodology, developmental trajectory and behavioral association. Chinese Science Bulletin, 2012, 57(35): 3399−3413 doi: 10.1360/972012-702 [11] Biswal B B, Mennes M, Zuo X N, Gohel S, Kelly C, Smith S M, et al. Toward discovery science of human brain function. Proceedings of the National Academy of Sciences of the United States of America, 2010, 107(10): 4734−4739 doi: 10.1073/pnas.0911855107 [12] Friston K J. Modalities, modes, and models in functional neuroimaging. Science, 2009, 326(5951): 399−403 doi: 10.1126/science.1174521 [13] Turk-Browne N B. Functional interactions as big data in the human brain. Science, 2013, 342(6158): 580−584 doi: 10.1126/science.1238409 [14] Park H J, Friston K. Structural and functional brain networks: From connections to cognition. Science, 2013, 342(6158): 1238411 doi: 10.1126/science.1238411 [15] Hein G, Morishima Y, Leiberg S, Sul S, Fehr E. The brain' s functional network architecture reveals human motives. Science, 2016, 351(6277): 1074−1078 doi: 10.1126/science.aac7992 [16] Tavor I, Jones O P, Mars R B, Smith S M, Behrens T E, Jbabdi S. Task-free MRI predicts individual differences in brain activity during task performance. Science, 2016, 352(6282): 216−220 doi: 10.1126/science.aad8127 [17] Field G D, Gauthier J L, Sher A, Greschner M, Machado T A, Jepson L H, et al. Functional connectivity in the retina at the resolution of photoreceptors. Nature, 2010, 467(7316): 673−677 doi: 10.1038/nature09424 [18] Honey C J, Sporns O, Cammoun L, Gigandet X, Thiran J P, Meuli R, et al. Predicting human resting-state functional connectivity from structural connectivity. Proceedings of the National Academy of Sciences of the United States of America, 2009, 106(6): 2035−2040 doi: 10.1073/pnas.0811168106 [19] Smith S M, Fox P T, Miller K L, Glahn D C, Fox M, Mackay C E, et al. Correspondence of the brain' s functional architecture during activation and rest. Proceedings of the National Academy of Sciences of the United States of America, 2009, 106(31): 13040−13045 doi: 10.1073/pnas.0905267106 [20] Riedl V, Utz L, Castrillon G, Grimmer T, Rauschecker J P, Ploner M, et al. Metabolic connectivity mapping reveals effective connectivity in the resting human brain. Proceedings of the National Academy of Sciences of the United States of America, 2016, 113(2): 428−433 doi: 10.1073/pnas.1513752113 [21] Hermundstad A M, Bassett D S, Brown K S, Aminoff E M, Clewett D, Freeman S, et al. Structural foundations of resting-state and task-based functional connectivity in the human brain. Proceedings of the National Academy of Sciences of the United States of America, 2013, 110(15): 6169−6174 doi: 10.1073/pnas.1219562110 [22] Buijink A W G, Van Der Stouwe A M M, Broersma M, Sharifi S, Groot P F C, Speelman J D, et al. Motor network disruption in essential tremor: A functional and effective connectivity study. Brain, 2015, 138(10): 2934−2947 doi: 10.1093/brain/awv225 [23] Gibson W S, Jo H J, Testini P, Cho S, Felmlee J P, Welker K M, et al. Functional correlates of the therapeutic and adverse effects evoked by thalamic stimulation for essential tremor. Brain, 2016, 139(8): 2198−2210 doi: 10.1093/brain/aww145 [24] Huang S, Li J, Ye J P, Fleisher A, Chen K W, Wu T, et al. A sparse structure learning algorithm for Gaussian Bayesian network identification from high-dimensional data. IEEE Transactions on Pattern Analysis and Machine Intelligence, 2013, 35(6): 1328−1342 doi: 10.1109/TPAMI.2012.129 [25] Zhou L P, Wang L, Liu L Q, Ogunbona P, Shen D G. Discriminative brain effective connectivity analysis for Alzheimer' s disease: A kernel learning approach upon sparse Gaussian Bayesian network. In: Proceedings of the 2013 IEEE Conference on Computer Vision and Pattern Recognition. Portland, USA: IEEE, 2013. 2243−2250 [26] Zhou L P, Wang L, Liu L Q, Ogunbona P, Shen D G. Learning discriminative Bayesian networks from high-dimensional continuous neuroimaging data. IEEE Transactions on Pattern Analysis and Machine Intelligence, 2016, 38(11): 2269−2283 doi: 10.1109/TPAMI.2015.2511754 [27] Huang S, Li J, Ye J P, Fleisher A, Chen K W, Wu T, et al. Brain effective connectivity modeling for Alzheimer’s disease by sparse Gaussian Bayesian network. In: Proceedings of the 17th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. San Diego, USA: ACM, 2011. 931−939 [28] Logothetis N K, Pauls J, Augath M, Trinath T, Oeltermann A. Neurophysiological investigation of the basis of the fMRI signal. Nature, 2001, 412(6843): 150−157 doi: 10.1038/35084005 [29] 宫殿坤. 人脑认知加工的时空连续性与可塑性研究[博士学位论文], 电子科技大学, 中国, 2015Gong Dian-Kun. Research on the temporal-spatial continuity and plasticity of cognitive processing of human brain [Ph. D. dissertation], University of Electronic Science and Technology of China, China, 2015 [30] Cox R W, Hyde J S. Software tools for analysis and visualization of fMRI data. NMR in Biomedicine, 1997, 10(4-5): 171−178 doi: 10.1002/(SICI)1099-1492(199706/08)10:4/5<171::AID-NBM453>3.0.CO;2-L [31] Avants B B, Tustison N J, Song G, Cook P A, Klein A, Gee J C. A reproducible evaluation of ANTs similarity metric performance in brain image registration. NeuroImage, 2011, 54(3): 2033−2044 doi: 10.1016/j.neuroimage.2010.09.025 [32] Jenkinson M, Beckmann C F, Behrens T E J, Woolrich M W, Smith S M. FSL. NeuroImage, 2012, 62(2): 782−790 doi: 10.1016/j.neuroimage.2011.09.015 [33] Friston K J, Ashburner J T, Kiebel S J, Nichols T E, Penny W D. Statistical Parametric Mapping: The Analysis of Functional Brain Images. Amsterdam: Academic Press, 2007. [34] Song X W, Dong Z Y, Long X Y, Li S F, Zuo X N, Zhu C Z, et al. REST: A toolkit for resting-state functional magnetic resonance imaging data processing. PLoS One, 2011, 6(9): e25031 doi: 10.1371/journal.pone.0025031 [35] Yan C G, Wang X D, Zuo X N, Zang Y F. DPABI: Data processing & analysis for (resting-state) brain imaging. Neuroinformatics, 2016, 14(3): 339−351 doi: 10.1007/s12021-016-9299-4 [36] Esteban O, Markiewicz C J, Blair R W, Moodie C A, Isik A I, Erramuzpe A, et al. fMRIPrep: A robust preprocessing pipeline for functional MRI. Nature Methods, 2019, 16(1): 111−116 doi: 10.1038/s41592-018-0235-4 [37] Gao Q, Duan X J, Chen H F. Evaluation of effective connectivity of motor areas during motor imagery and execution using conditional Granger causality. NeuroImage, 2011, 54(2): 1280−1288 doi: 10.1016/j.neuroimage.2010.08.071 [38] Zuo X N, Kelly C, Adelstein J S, Klein D F, Castellanos F X, Milham M P. Reliable intrinsic connectivity networks: Test-retest evaluation using ICA and dual regression approach. NeuroImage, 2010, 49(3): 2163−2177 doi: 10.1016/j.neuroimage.2009.10.080 [39] Liu C H, JaJa J, Pessoa L. LEICA: Laplacian eigenmaps for group ICA decomposition of fMRI data. NeuroImage, 2018, 169: 363−373 doi: 10.1016/j.neuroimage.2017.12.018 [40] Tzourio-Mazoyer N, Landeau B, Papathanassiou D, Crivello F, Etard O, Delcroix N, et al. Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. NeuroImage, 2002, 15(1): 273−289 doi: 10.1006/nimg.2001.0978 [41] Shattuck D W, Mirza M, Adisetiyo V, Hojatkashani C, Salamon G, Narr K L, et al. Construction of a 3D probabilistic atlas of human cortical structures. NeuroImage, 2008, 39(3): 1064−1080 doi: 10.1016/j.neuroimage.2007.09.031 [42] Fan L Z, Li H, Zhuo J J, Zhang Y, Wang J J, Chen L F, et al. The human brainnetome atlas: A new brain atlas based on connectional architecture. Cerebral Cortex, 2016, 26(8): 3508−3526 doi: 10.1093/cercor/bhw157 [43] Glasser M F, Coalson T S, Robinson E C, Hacker C D, Harwell J, Yacoub E, et al. A multi-modal parcellation of human cerebral cortex. Nature, 2016, 536(7615): 171−178 doi: 10.1038/nature18933 [44] Ji J Z, Liu J D, Liang P P, Zhang A D. Learning effective connectivity network structure from fMRI data based on artificial immune algorithm. PLoS One, 2016, 11(4): e0152600 doi: 10.1371/journal.pone.0152600 [45] Liu J D, Ji J Z, Zhang A D, Liang P P. An ant colony optimization algorithm for learning brain effective connectivity network from fMRI data. In: Proceedings of the 2016 IEEE International Conference on Bioinformatics and Biomedicine. Shenzhen, China: IEEE, 2016. 360−367 [46] Park H J, Friston K J, Pae C, Park B, Razi A. Dynamic effective connectivity in resting state fMRI. NeuroImage, 2018, 180: 594−608 doi: 10.1016/j.neuroimage.2017.11.033 [47] Zhou Z Y, Chen Y H, Ding M Z, Wright P, Lu Z H, Liu Y J. Analyzing brain networks with PCA and conditional Granger causality. Human Brain Mapping, 2009, 30(7): 2197−2206 doi: 10.1002/hbm.20661 [48] Sanchez-Romero R, Ramsey J D, Zhang K, Glymour M R K, Huang B W, Glymour C. Estimating feedforward and feedback effective connections from fMRI time series: Assessments of statistical methods. Network Neuroscience, 2019, 3(2): 274−306 doi: 10.1162/netn_a_00061 [49] McLntosh A R, Gonzalez-Lima F. Structural equation modeling and its application to network analysis in functional brain imaging. Human Brain Mapping, 1994, 2(1−2): 2−22 doi: 10.1002/hbm.460020104 [50] Spirtes P, Glymour C, Scheines R. Causation, Prediction, and Search (Second edition). Cambridge: MIT Press, 2000. [51] Ramnani N, Behrens T E J, Penny W, Matthews P M. New approaches for exploring anatomical and functional connectivity in the human brain. Biological Psychiatry, 2004, 56(9): 613−619 doi: 10.1016/j.biopsych.2004.02.004 [52] Penny W D, Stephan K E, Mechelli A, Friston K J. Modelling functional integration: A comparison of structural equation and dynamic causal models. NeuroImage, 2004, 23(Suppl 1): S264−S274 [53] Shimizu S, Hoyer P O, Hyvarinen A, Kerminen A. A linear non-Gaussian acyclic model for causal discovery. Journal of Machine Learning Research, 2006, 7: 2003−2030 [54] Shimizu S, Inazumi T, Sogawa Y, Hyvarinen A, Kerminen A, Washio T, et al. DirectLiNGAM: A direct method for learning a linear non-Gaussian structural equation model. Journal of Machine Learning Research, 2011, 12: 1225−1248 [55] Kim J, Zhu W, Chang L D, Bentler P M, Ernst T. Unified structural equation modeling approach for the analysis of multisubject, multivariate functional MRI data. Human Brain Mapping, 2007, 28(2): 85−93 doi: 10.1002/hbm.20259 [56] Gates K M, Molenaar P C M, Hillary F G, Slobounov S. Extended unified SEM approach for modeling event-related fMRI data. NeuroImage, 2011, 54(2): 1151−1158 doi: 10.1016/j.neuroimage.2010.08.051 [57] Hillary F G, Medaglia J D, Gates K, Molenaar P C, Slocomb J, Peechatka A, et al. Examining working memory task acquisition in a disrupted neural network. Brain, 2011, 134(5): 1555−1570 doi: 10.1093/brain/awr043 [58] Gates K M, Molenaar P C M. Group search algorithm recovers effective connectivity maps for individuals in homogeneous and heterogeneous samples. NeuroImage, 2012, 63(1): 310−319 doi: 10.1016/j.neuroimage.2012.06.026 [59] Henry T R, Feczko E, Cordova M, Earl E, Williams S, Nigg J T, et al. Comparing directed functional connectivity between groups with confirmatory subgrouping GIMME. NeuroImage, 2019, 188: 642−653 doi: 10.1016/j.neuroimage.2018.12.040 [60] 姚垚, 冀俊忠. 基于栈式循环神经网络的血液动力学状态估计方法. 自动化学报, 2020, 46(5): 991−1003Yao Yao, Ji Jun-Zhong. Estimation of hemodynamic states based on stacked recurrent neural network in fMRI time series. Acta Automatica Sinica, 2020, 46(5): 991−1003 [61] Li B J, Daunizeau J, Stephan K E, Penny W, Hu D W, Friston K. Generalised filtering and stochastic DCM for fMRI. NeuroImage, 2011, 58(2): 442−457 doi: 10.1016/j.neuroimage.2011.01.085 [62] Friston K J, Kahan J, Biswal B, Razi A. A DCM for resting state fMRI. NeuroImage, 2014, 94: 396−407 doi: 10.1016/j.neuroimage.2013.12.009 [63] Razi A, Kahan J, Rees G, Friston K J. Construct validation of a DCM for resting state fMRI. NeuroImage, 2015, 106: 1−14 doi: 10.1016/j.neuroimage.2014.11.027 [64] Daunizeau J, David O, Stephan K E. Dynamic causal modelling: A critical review of the biophysical and statistical foundations. NeuroImage, 2011, 58(2): 312−322 doi: 10.1016/j.neuroimage.2009.11.062 [65] Seghier M L, Friston K J. Network discovery with large DCMs. NeuroImage, 2013, 68: 181−191 doi: 10.1016/j.neuroimage.2012.12.005 [66] Razi A, Seghier M L, Zhou Y, McColgan P, Zeidman P, Park H J, et al. Large-scale DCMs for resting-state fMRI. Network Neuroscience, 2017, 1(3): 222−241 doi: 10.1162/NETN_a_00015 [67] Frassle S, Lomakina E I, Razi A, Friston K J, Buhmann J M, Stephan K E. Regression DCM for fMRI. NeuroImage, 2017, 155: 406−421 doi: 10.1016/j.neuroimage.2017.02.090 [68] Frassle S, Lomakina E I, Kasper L, Manjaly Z M, Leff A, Pruessmann K P, et al. A generative model of whole-brain effective connectivity. NeuroImage, 2018, 179: 505−529 doi: 10.1016/j.neuroimage.2018.05.058 [69] Friston K, Zeidman P, Litvak V. Empirical Bayes for DCM: A group inversion scheme. Frontiers in Systems Neuroscience, 2015, 9: 164 [70] Friston K J, Litvak V, Oswal A, Razi A, Stephan K E, Van Wijk B C M, et al. Bayesian model reduction and empirical Bayes for group (DCM) studies. NeuroImage, 2016, 128: 413−431 doi: 10.1016/j.neuroimage.2015.11.015 [71] Gilson M, Moreno-Bote R, Ponce-Alvarez A, Ritter P, Deco G. Estimation of directed effective connectivity from fMRI functional connectivity hints at asymmetries of cortical connectome. PLoS Computational Biology, 2016, 12(3): e1004762 doi: 10.1371/journal.pcbi.1004762 [72] Gilson M, Deco G, Friston K J, Hagmann P, Mantini D, Betti V, et al. Effective connectivity inferred from fMRI transition dynamics during movie viewing points to a balanced reconfiguration of cortical interactions. NeuroImage, 2018, 180: 534−546 doi: 10.1016/j.neuroimage.2017.09.061 [73] Schiefer J, Niederbuhl A, Pernice V, Lennartz C, Hennig J, LeVan P, et al. From correlation to causation: Estimating effective connectivity from zero-lag covariances of brain signals. PLoS Computational Biology, 2018, 14(3): e1006056 doi: 10.1371/journal.pcbi.1006056 [74] Ramsey J D, Hanson S J, Hanson C, Halchenko Y O, Poldrack R A, Glymour C. Six problems for causal inference from fMRI. NeuroImage, 2010, 49(2): 1545−1558 doi: 10.1016/j.neuroimage.2009.08.065 [75] Li R, Chen K W, Zhang N, Fleisher A S, Yao L, Wu X. Effective connectivity analysis of default mode network based on the Bayesian network learning approach. In: Proceedings of the Medical Imaging 2009: Biomedical Applications in Molecular, Structural, and Functional Imaging. Lake Buena Vista (Orlando Area), USA: SPIE, 2009. Article No. 72621W [76] Zheng X B, Rajapakse J C. Learning functional structure from fMR images. NeuroImage, 2006, 31(4): 1601−1613 doi: 10.1016/j.neuroimage.2006.01.031 [77] Rajapakse J C, Zhou J. Learning effective brain connectivity with dynamic Bayesian networks. NeuroImage, 2007, 37(3): 749−760 doi: 10.1016/j.neuroimage.2007.06.003 [78] Burge J, Lane T, Link H, Qiu S B, Clark V P. Discrete dynamic Bayesian network analysis of fMRI data. Human Brain Mapping, 2009, 30(1): 122−137 doi: 10.1002/hbm.20490 [79] 冀俊忠, 刘金铎, 邹爱笑, 杨翠翠. 一种融合多源信息的脑效应连接网络蚁群学习算法. 自动化学报, DOI: 10.16383/j.aas.c180680. 2019Ji Jun-Zhong, Liu Jin-Duo, Zou Ai-Xiao, Yang Cui-Cui. An ant colony optimization algorithm merged with multiple source information for learning brain effective connectivity networks. Acta Automatica Sinica, DOI: 10.16383/j.aas.c180680. 2019. [80] Wu X, Wen X Y, Li J, Yao L. A new dynamic Bayesian network approach for determining effective connectivity from fMRI data. Neural Computing and Applications, 2014, 24(1): 91−97 doi: 10.1007/s00521-013-1465-0 [81] Dang S, Chaudhury S, Lall B, Roy P K. The dynamic programming high-order dynamic Bayesian networks learning for identifying effective connectivity in human brain from fMRI. Journal of Neuroscience Methods, 2017, 285: 33−44 doi: 10.1016/j.jneumeth.2017.05.009 [82] Dang S, Chaudhury S, Lall B, Roy P K. Tractography-based score for learning effective connectivity from multimodal imaging data using dynamic Bayesian networks. IEEE Transactions on Biomedical Engineering, 2018, 65(5): 1057−1068 [83] Harrison L, Penny W D, Friston K J. Multivariate autoregressive modeling of fMRI time series. NeuroImage, 2003, 19(4): 1477−1491 doi: 10.1016/S1053-8119(03)00160-5 [84] Valdes-Sosa P A, Roebroeck A, Daunizeau J, Friston K. Effective connectivity: Influence, causality and biophysical modeling. NeuroImage, 2011, 58(2): 339−361 doi: 10.1016/j.neuroimage.2011.03.058 [85] Liao W, Mantini D, Zhang Z Q, Pan Z Y, Ding J R, Gong Q Y, et al. Evaluating the effective connectivity of resting state networks using conditional Granger causality. Biological Cybernetics, 2010, 102(1): 57−69 doi: 10.1007/s00422-009-0350-5 [86] Guo S X, Seth A K, Kendrick K M, Zou C, Feng J F. Partial Granger causality—eliminating exogenous inputs and latent variables. Journal of Neuroscience Methods, 2008, 172(1): 79−93 doi: 10.1016/j.jneumeth.2008.04.011 [87] Baccala L A, Sameshima K. Partial directed coherence: A new concept in neural structure determination. Biological Cybernetics, 2001, 84(6): 463−474 doi: 10.1007/PL00007990 [88] Eichler M. On the Evaluation of Information flow in multivariate systems by the directed transfer function. Biological Cybernetics, 2006, 94(6): 469−482 doi: 10.1007/s00422-006-0062-z [89] Wu M H, Frye R E, Zouridakis G. A comparison of multivariate causality based measures of effective connectivity. Computers in Biology and Medicine, 2011, 41(12): 1132−1141 doi: 10.1016/j.compbiomed.2011.06.007 [90] Karanikolas G V, Giannakis G B. Identifying directional connections in brain networks via multi-kernel granger models. In: Proceedings of the 2017 IEEE International Conference on Acoustics, Speech and Signal Processing. New Orleans, USA: IEEE, 2017. 6304−6308 [91] Lobier M, Siebenhuhner F, Palva S, Palva J M. Phase transfer entropy: A novel phase-based measure for directed connectivity in networks coupled by oscillatory interactions. NeuroImage, 2014, 85: 853−872 doi: 10.1016/j.neuroimage.2013.08.056 [92] Ting C M, Seghouane A K, Salleh S H, Noor A M. Estimating effective connectivity from fMRI data using factor-based subspace autoregressive models. IEEE Signal Processing Letters, 2015, 22(6): 757−761 doi: 10.1109/LSP.2014.2365634 [93] Samdin S B, Ting C M, Ombao H, Salleh S H. A unified estimation framework for state-related changes in effective brain connectivity. IEEE Transactions on Biomedical Engineering, 2017, 64(4): 844−858 doi: 10.1109/TBME.2016.2580738 [94] Ting C M, Ombao H, Samdin S B, Salleh S H. Estimating dynamic connectivity states in fMRI using regime-switching factor models. IEEE Transactions on Medical Imaging, 2018, 37(4): 1011−1023 doi: 10.1109/TMI.2017.2780185 [95] Wei H L, An J, Shen H, Zeng L L, Qiu S J, Hu D W. Altered effective connectivity among core neurocognitive networks in idiopathic generalized epilepsy: An fMRI evidence. Frontiers in Human Neuroscience, 2016, 10: 447 [96] Meier J, Zhou X, Hillebrand A, Tewarie P, Stam C J, Van Mieghem P. The epidemic spreading model and the direction of information flow in brain networks. NeuroImage, 2017, 152: 639−646 doi: 10.1016/j.neuroimage.2017.02.007 [97] Chockanathan U, DSouza A M, Abidin A Z, Schifitto G, Wismuller A. Automated diagnosis of HIV-associated neurocognitive disorders using large-scale Granger causality analysis of resting-state functional MRI. Computers in Biology and Medicine, 2019, 106: 24−30 doi: 10.1016/j.compbiomed.2019.01.006 [98] Ambrogioni L, Hinne M, Van Gerven M A J, Maris E. GP CaKe: Effective brain connectivity with causal kernels. In: Proceedings of the 31st International Conference on Neural Information Processing Systems. Long Beach, USA: ACM, 2017. 951−960 [99] Farokhzadi M, Hossein-Zadeh G A, Soltanian-Zadeh H. Nonlinear effective connectivity measure based on adaptive Neuro Fuzzy Inference System and Granger Causality. NeuroImage, 2018, 181: 382−394 doi: 10.1016/j.neuroimage.2018.07.024 [100] Xu N, Spreng R N, Doerschuk P C. Initial validation for the estimation of resting-state fMRI effective connectivity by a generalization of the correlation approach. Frontiers in Neuroscience, 2017, 11: 271 doi: 10.3389/fnins.2017.00271 [101] Quian Quiroga R, Kraskov A, Kreuz T, Grassberger P. Performance of different synchronization measures in real data: A case study on electroencephalographic signals. Physical Review E, 2002, 65(4): 41903 doi: 10.1103/PhysRevE.65.041903 [102] Dsouza A M, Abidin A Z, Chockanathan U, Schifitto G, Wismuller A. Mutual connectivity analysis of resting-state functional MRI data with local models. NeuroImage, 2018, 178: 210−223 doi: 10.1016/j.neuroimage.2018.05.038 [103] Patel R S, Bowman D B, Rilling J K. A Bayesian approach to determining connectivity of the human brain. Human Brain Mapping, 2006, 27(3): 267−276 doi: 10.1002/hbm.20182 [104] Hyvärinen A, Smith S M. Pairwise likelihood ratios for estimation of non-Gaussian structural equation models. Journal of Machine Learning Research, 2013, 14(1): 111−152 [105] Bielczyk N Z, Llera A, Buitelaar J K, Glennon J C, Beckmann C F. Increasing robustness of pairwise methods for effective connectivity in Magnetic Resonance Imaging by using fractional moment series of BOLD signal distributions. Network Neuroscience, 2019, 3(4): 1009−1037 doi: 10.1162/netn_a_00099 [106] Lee N, Kim J M. Copula directional dependence for inference and statistical analysis of whole-brain connectivity from fMRI data. Brain and Behavior, 2019, 9(1): e01191 doi: 10.1002/brb3.1191 [107] Chiang S, Guindani M, Yeh H J, Haneef Z, Stern J M, Vannucci M. Bayesian vector autoregressive model for multi-subject effective connectivity inference using multi-modal neuroimaging data. Human Brain Mapping, 2017, 38(3): 1311−1332 doi: 10.1002/hbm.23456 [108] Anwar A R, Muthalib M, Perrey S, Galka A, Granert O, Wolff S, et al. Effective connectivity of cortical sensorimotor networks during finger movement tasks: A simultaneous fNIRS, fMRI, EEG study. Brain Topography, 2016, 29(5): 645−660 doi: 10.1007/s10548-016-0507-1 [109] 纪子龙, 冀俊忠, 刘金铎, 杨翠翠. 基于萤火虫算法的脑效应连接网络学习方法. 哈尔滨工业大学学报, 2019, 51(5): 76−84 doi: 10.11918/j.issn.0367-6234.201811073Ji Zi-Long, Ji Jun-Zhong, Liu Jin-Duo, Yang Cui-Cui. Learning effective connectivity network structure based on firefly algorithm. Journal of Harbin Institute of Technology, 2019, 51(5): 76−84 doi: 10.11918/j.issn.0367-6234.201811073 [110] 纪子龙, 冀俊忠. 基于双萤火虫种群并行搜索的脑效应连接网络学习方法. 浙江大学学报(工学版), 2020, 54(4): 694−703, 777Ji Zi-Long, Ji Jun-Zhong. Learning effective connectivity network structure based on parallel searching of double firefly populations. Journal of Zhejiang University (Engineering Science), 2020, 54(4): 694−703, 777 [111] Goudet O, Kalainathan D, Caillou P, Guyon I, Lopez-Paz D, Sebag M. Causal generative neural networks. arXiv: 1711.08936v2, 2018. [112] Kalainathan D, Goudet O, Guyon I, Lopez-Paz D, Sebag M. SAM: Structural agnostic model, causal discovery and penalized adversarial learning. arXiv: 1803.04929, 2018. [113] Njah H, Jamoussi S, Mahdi W. Deep Bayesian network architecture for Big Data mining. Concurrency and Computation Practice and Experience, 2019, 31(2): e4418 doi: 10.1002/cpe.4418 [114] Dasgupta I, Wang J, Chiappa S, Mitrovic J, Ortega P, Raposo D, et al. Causal reasoning from meta-reinforcement learning. arXiv: 1901.08162v1, 2019. [115] Faes L, Stramaglia S, Marinazzo D. On the interpretability and computational reliability of frequency-domain Granger causality. F1000Research, 2017, 6: 1710 doi: 10.12688/f1000research.12694.1 -



 下载:
下载: