Medical Small Lesion Segmentation Network Based on Physical Saliency Feature Perception
-
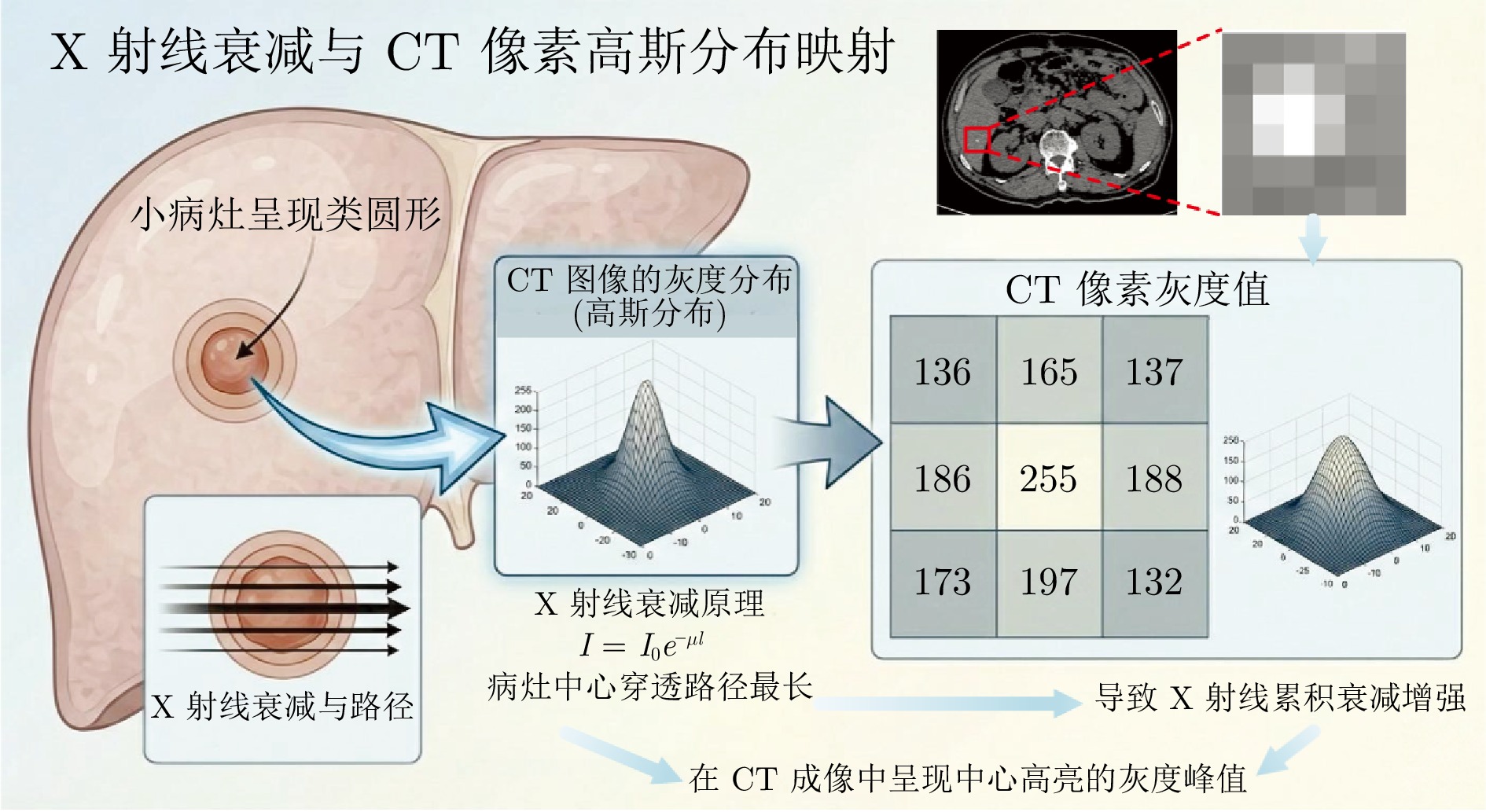
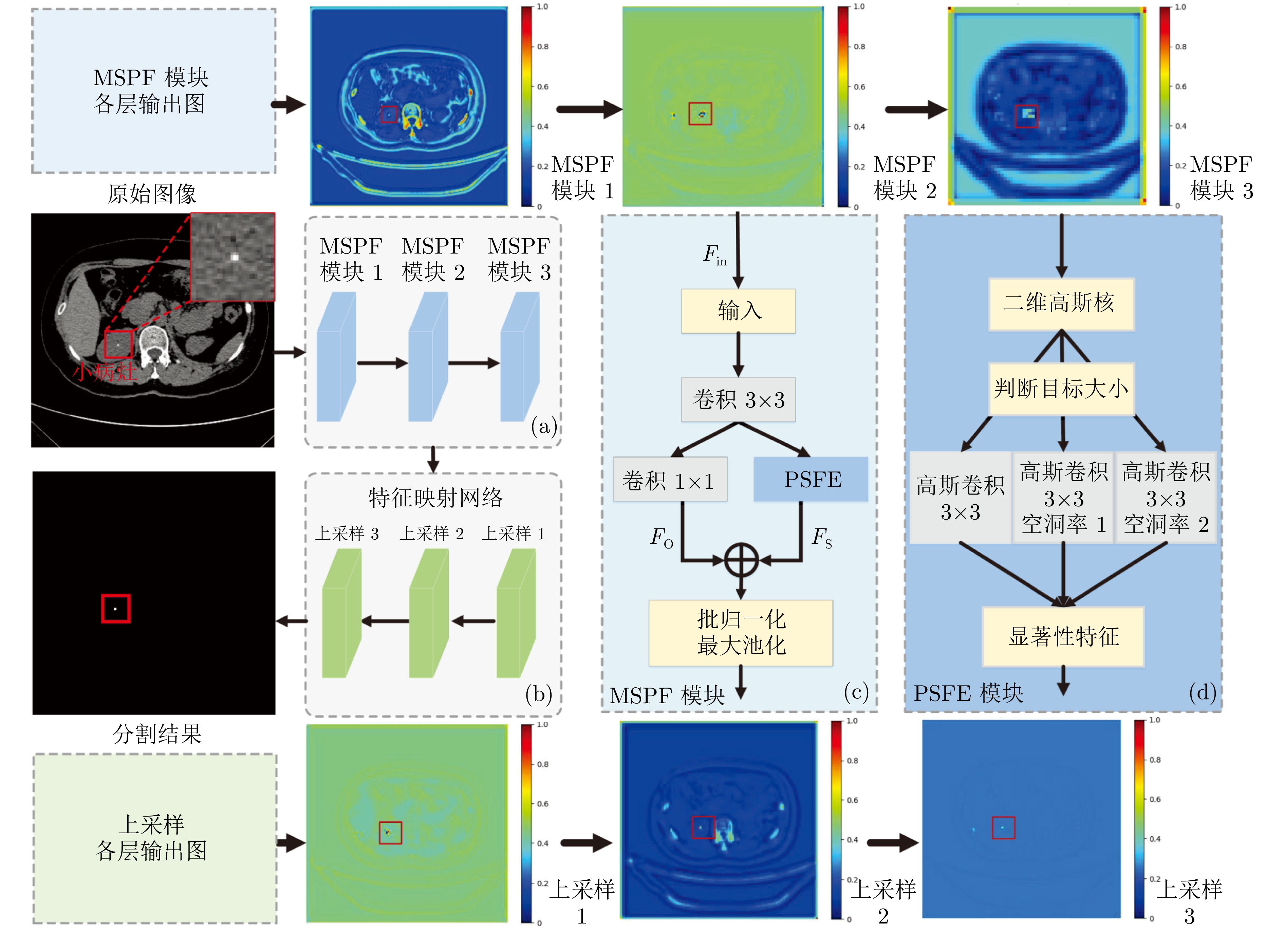
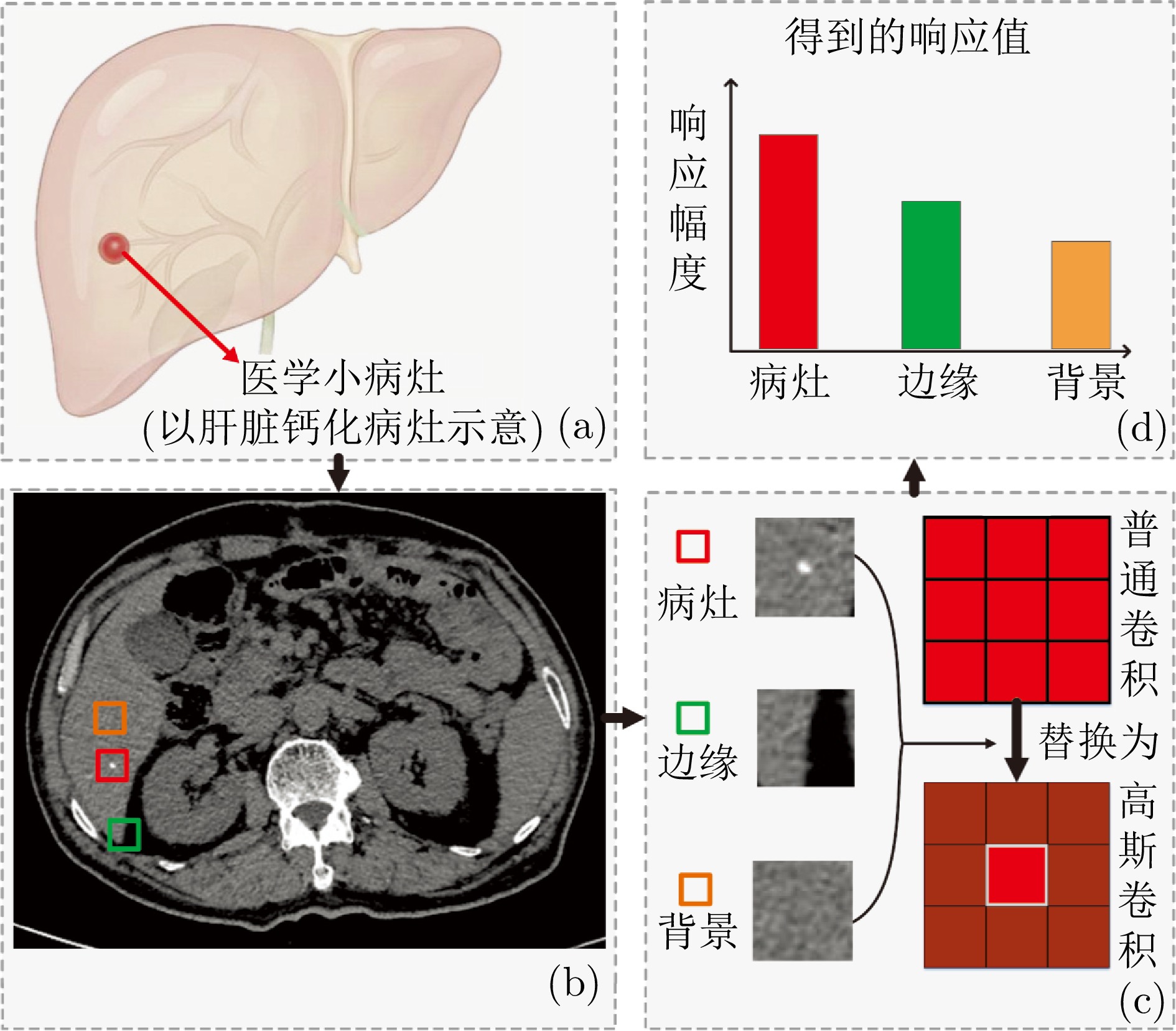
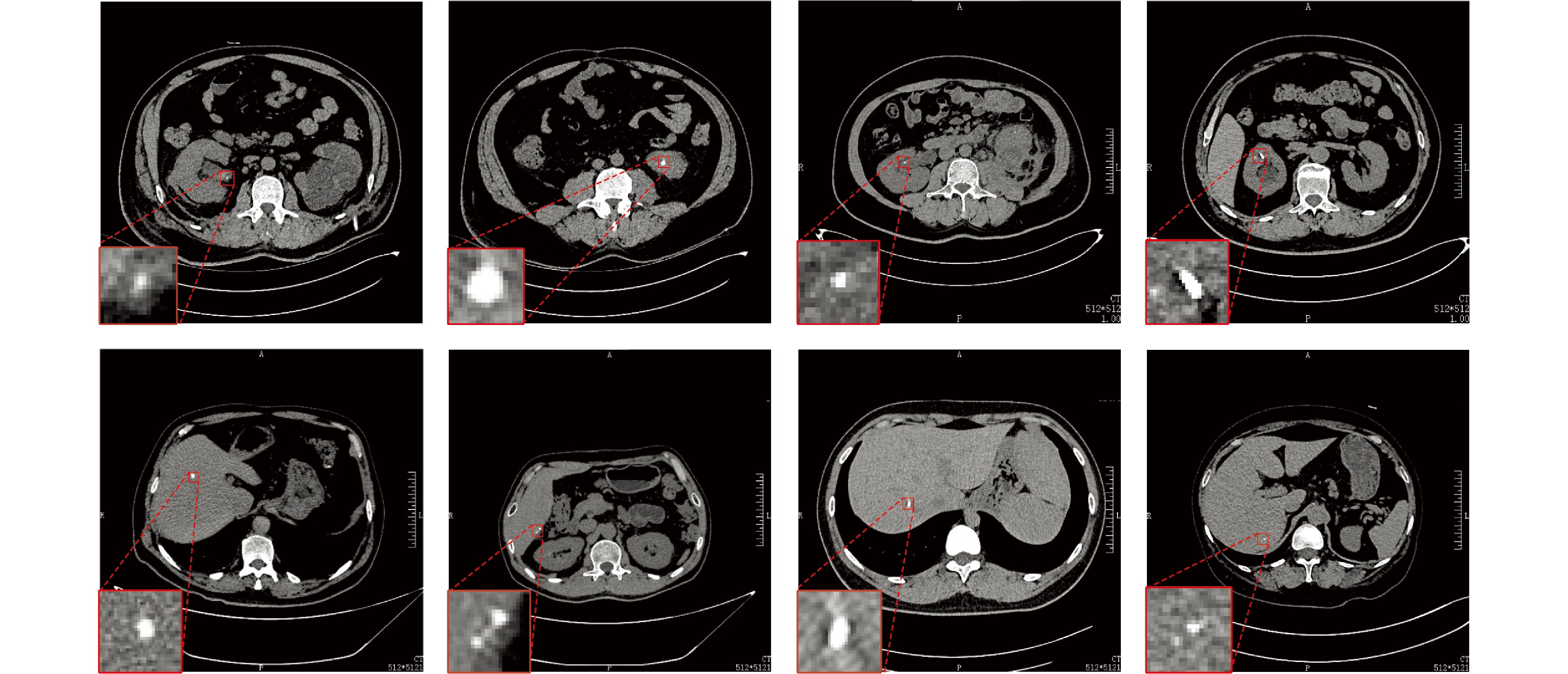
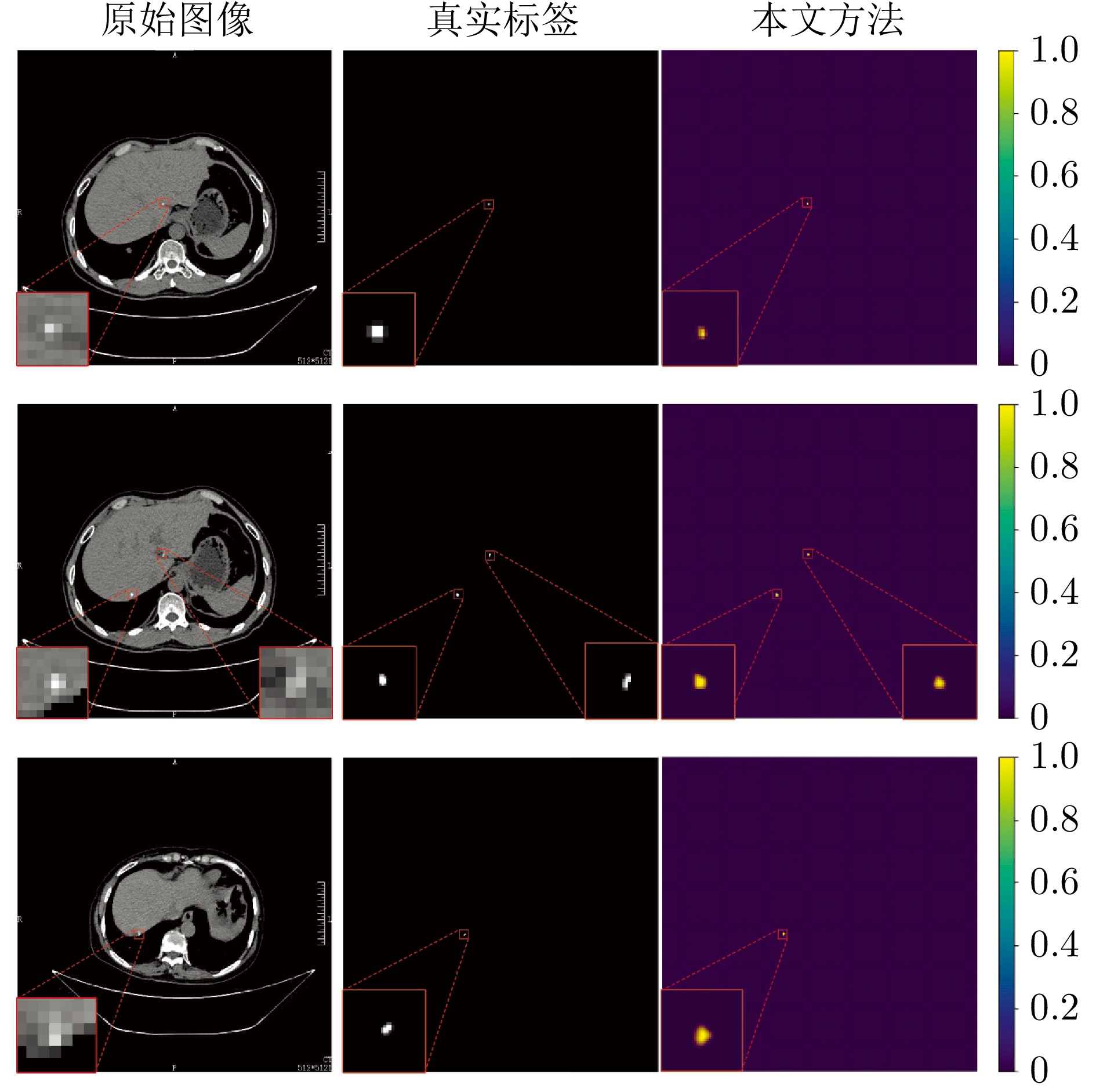
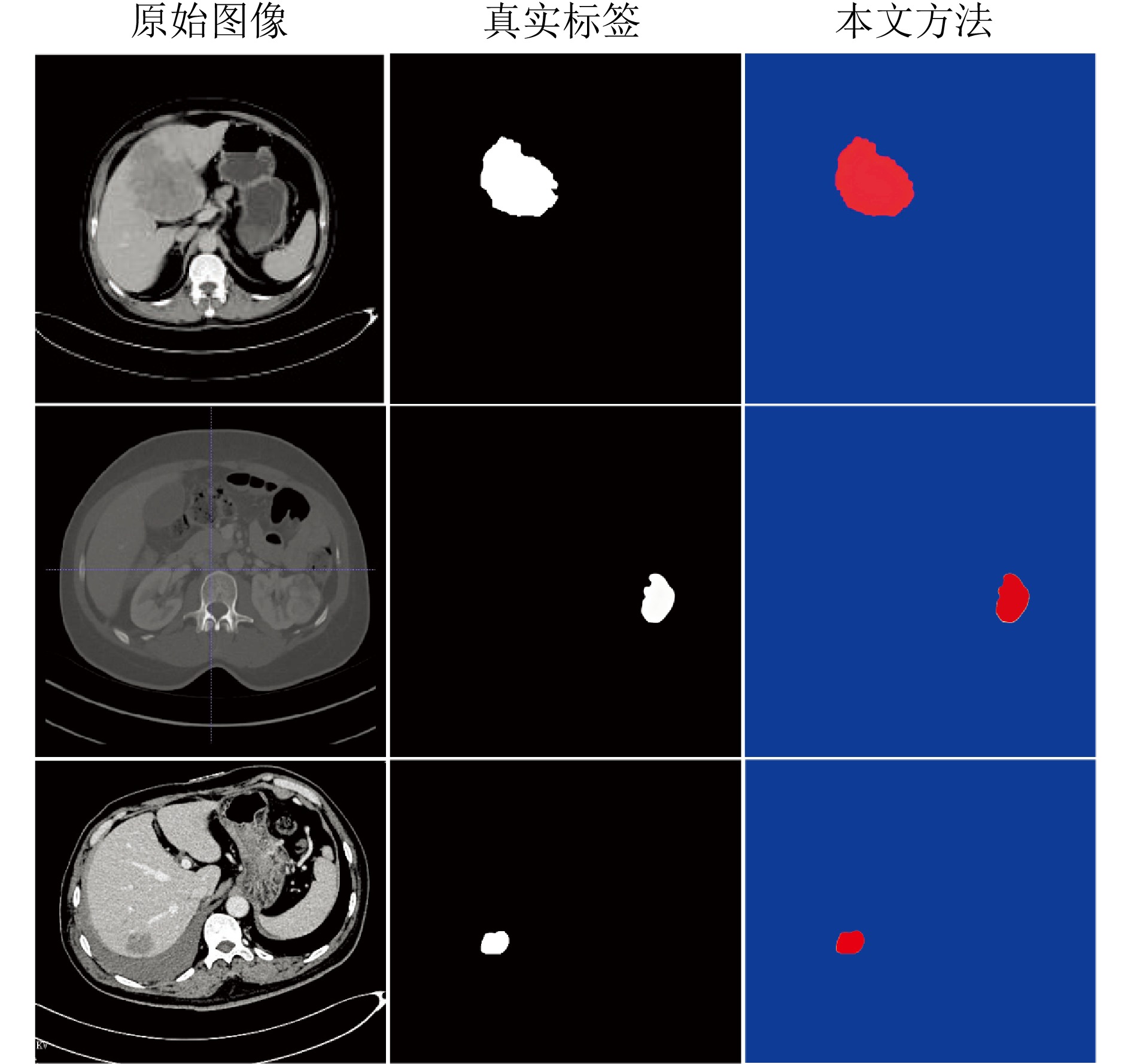
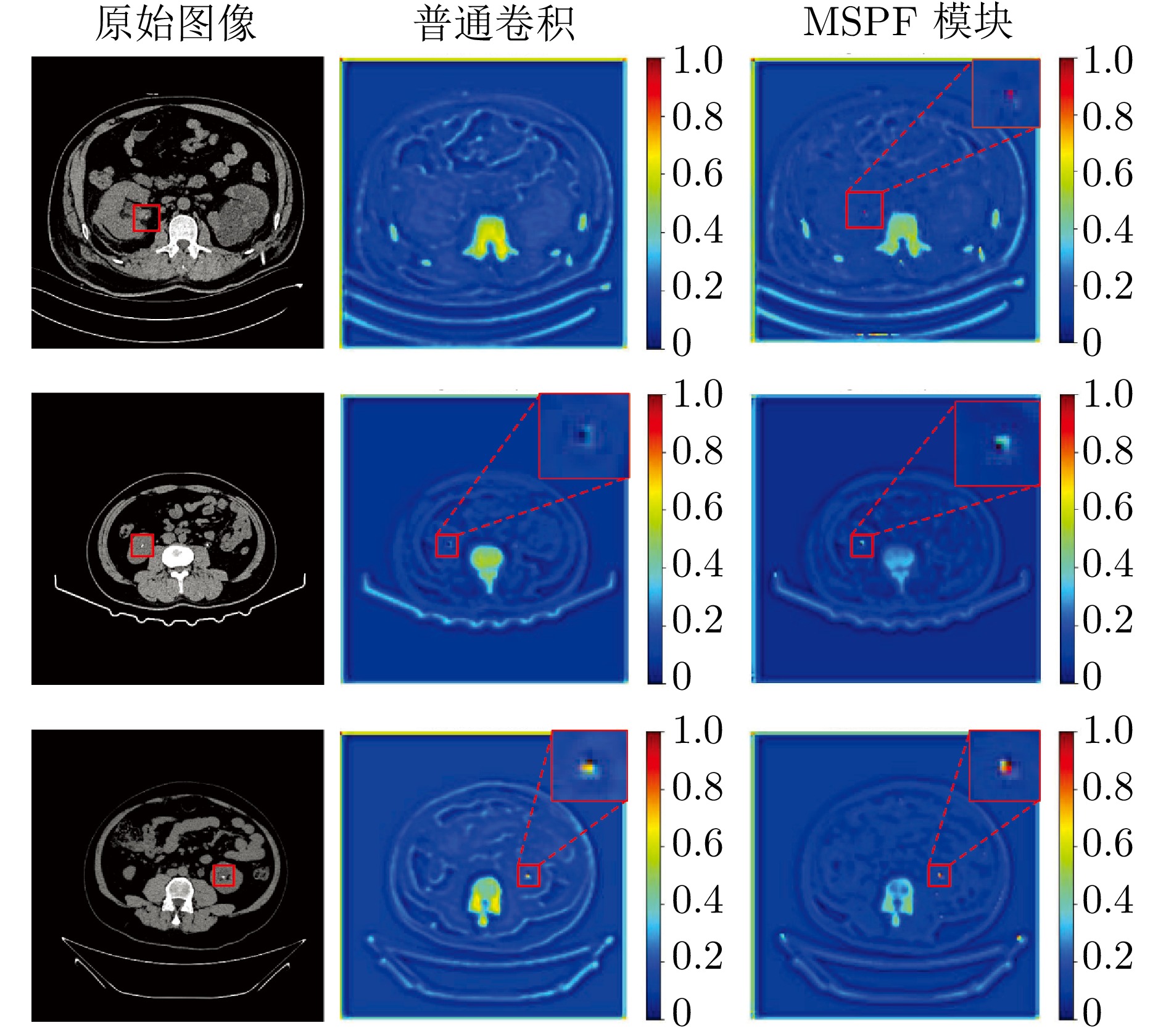
摘要: 精准分割早期小病灶对疾病诊疗至关重要, 但现有方法面对特征稀疏、易受背景干扰的小病灶时性能显著下降. 受计算机断层扫描成像中X射线衰减物理原理启发, 发现小病灶在影像中呈现出中心亮、边缘渐弱的强度分布, 其轮廓与二维高斯分布高度吻合. 为此, 提出一种辐射衰减原理引导的显著性感知分割网络(RAP-Net). RAP-Net将射线衰减导致的二维高斯分布作为相关性滤波卷积核融入深度学习架构, 并通过专为小病灶设计的多尺度感知特征网络, 其核心显著特征感知提取模块利用多尺度高斯空洞卷积精确建模衰减原理以实现稀疏特征的深度挖掘与背景抑制. 实验表明, RAP-Net在小肾结石与小肝脏钙化灶分割任务中的Dice系数和IoU提升至少18.05%与19.55%, 显著超越现有主流方法.Abstract: Accurate segmentation of early-stage small lesions is crucial for disease diagnosis and treatment. However, existing methods suffer from a significant drop in performance when dealing with small lesions that are feature-sparse and susceptible to background interference. Inspired by the physical principles of X-ray attenuation in computed tomography imaging, we observe that small lesions exhibit an intensity distribution characterized by a bright center and gradually fading edges, with their contours closely matching a two-dimensional Gaussian distribution. To address this, we propose a radiation attenuation principle-guided saliency perception segmentation network (RAP-Net). RAP-Net integrates the two-dimensional Gaussian distribution resulting from radiation attenuation as a correlation filtering convolution kernel into a deep learning architecture. Through a multi-scale perceptual feature network specifically designed for small lesions, its core perceptual salient feature extraction utilizes multi-scale Gaussian dilated convolution to accurately model the attenuation principle, thereby achieving deep mining of sparse features and background suppression. Experiments demonstrate that RAP-Net achieves at least 18.05% and 19.55% improvements in Dice coefficient and IoU, respectively, for the segmentation tasks of small kidney stones and small liver calcifications, significantly outperforming existing mainstream methods.
-
表 1 不同方法在小肾结石分割任务中的Dice系数与IoU
Table 1 The Dice coefficient and IoU of different methods in small kidney stone segmentation task
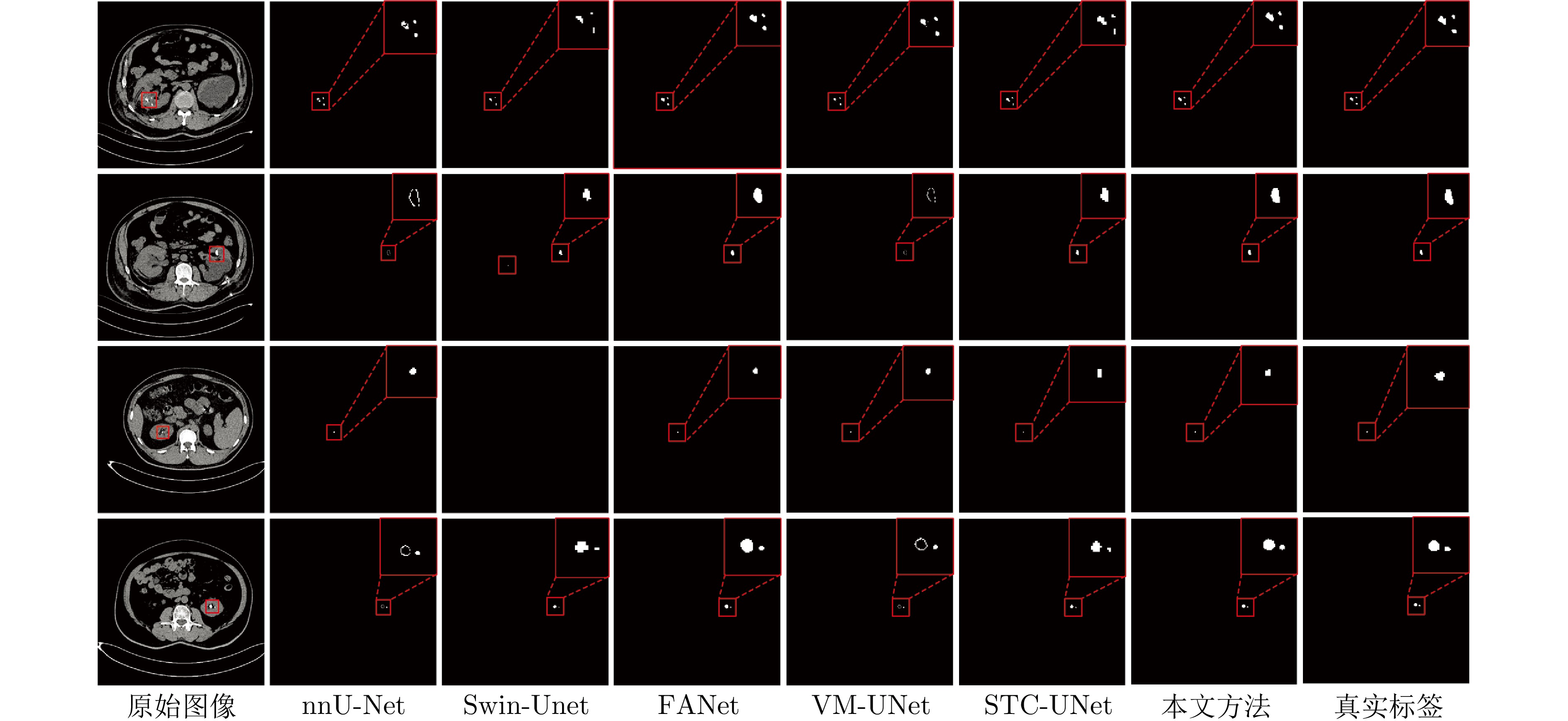
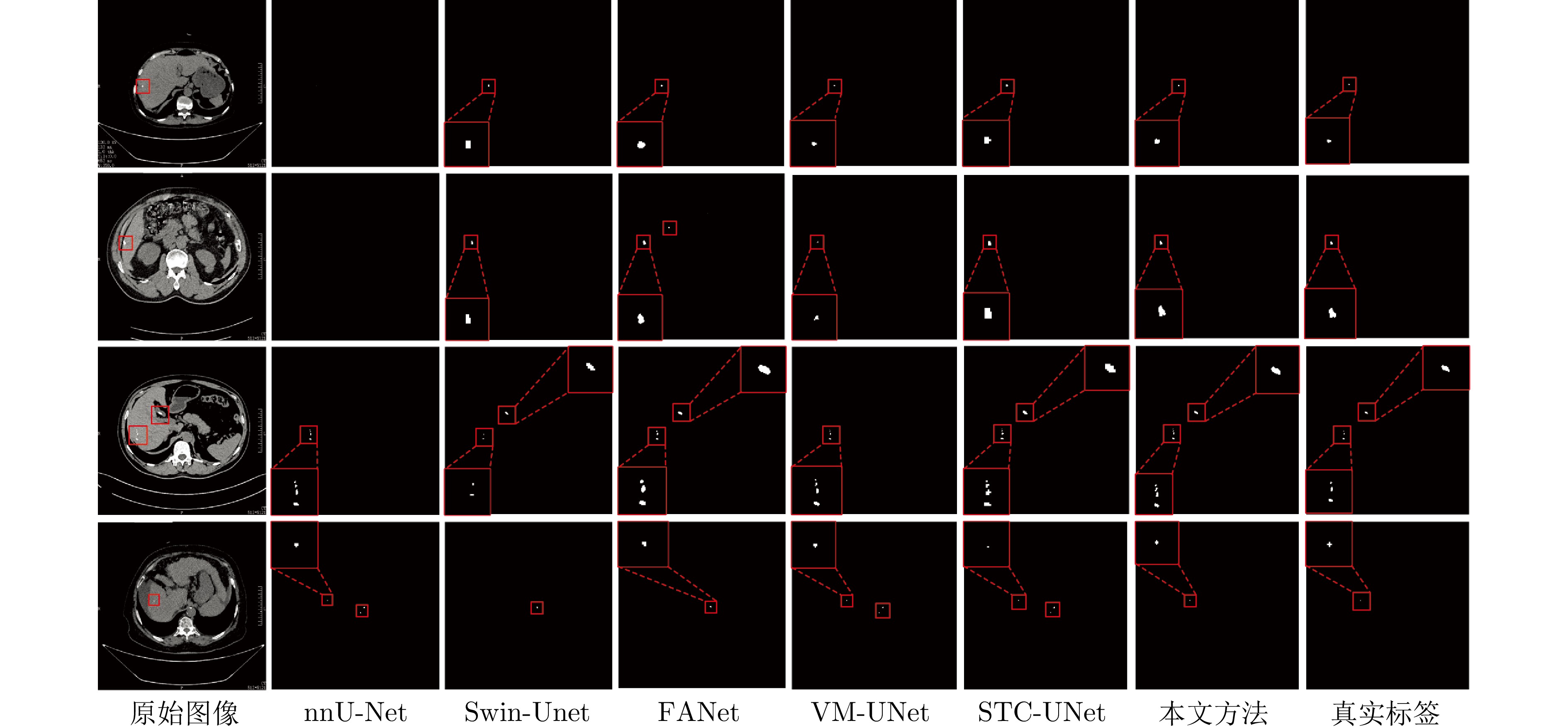
方法 Dice (%) IoU (%) 参数量(M) nnU-Net[30] 63.85 51.34 30.60 FANet[33] 61.35 46.19 34.15 Swin-Unet[36] 64.79 49.32 27.20 STC-UNet[37] 61.47 52.29 41.06 VM-UNet[38] 58.90 44.12 27.43 Zig-RiR[39] 65.48 55.15 24.58 MedSAM[40] 59.64 47.71 93.74 本文方法(3-layer MSPF) 86.44 77.91 0.34 本文方法(5-layer MSPF) 86.65 78.05 — 本文方法(7-layer MSPF) 87.43 78.51 — 表 2 不同方法在小肝脏钙化灶分割任务中的Dice系数与IoU
Table 2 The Dice coefficient and IoU of different methods in small liver calcification segmentation task
方法 Dice (%) IoU (%) 参数量(M) nnU-Net[30] 53.03 44.45 30.60 FANet[33] 44.12 35.70 34.15 Swin-Unet[36] 37.81 38.44 27.20 STC-UNet[37] 43.12 31.85 41.06 VM-UNet[38] 27.81 20.19 27.43 Zig-RiR[39] 66.59 56.02 24.58 MedSAM[40] 55.13 45.06 93.74 本文方法(3-layer MSPF) 84.64 75.57 0.34 本文方法(5-layer MSPF) 84.96 75.84 — 本文方法(7-layer MSPF) 85.15 76.02 — 表 3 小肾结石分割任务中消融实验的Dice系数与IoU
Table 3 The Dice coefficient and IoU of ablation experiment in small kidney stone segmentation task
-
[1] Zhang K J, Zhang L, Pan H W. UoloNet: Based on multi-tasking enhanced small target medical segmentation model. Artificial Intelligence Review, 2024, 57(2): Article No. 31 doi: 10.1007/s10462-023-10671-5 [2] Koleilat T, Asgariandehkordi H, Rivaz H, Xiao Y M. MedCLIP-SAMv2: Towards universal text-driven medical image segmentation. Medical Image Analysis, 2025, 106: Article No. 103749 doi: 10.1016/j.media.2025.103749 [3] He A L, Wang K, Li T, Du C K, Xia S, Fu H Z. H2Former: An efficient hierarchical hybrid transformer for medical image segmentation. IEEE Transactions on Medical Imaging, 2023, 42(9): 2763−2775 doi: 10.1109/TMI.2023.3264513 [4] Zhou X D, Chen T X. BSBP-RWKV: Background suppression with boundary preservation for efficient medical image segmentation. In: Proceedings of the 32nd ACM International Conference on Multimedia. Melbourne, Australia: Association for Computing Machinery, 2024. 4938-4946 [5] Mou L, Zhao Y T, Fu H Z, Liu Y H, Cheng J, Zheng Y L, et al. CS.2-Net: Deep learning segmentation of curvilinear structures in medical imaging. Medical Image Analysis, 2021, 67: Article No. 101874 doi: 10.1016/j.media.2020.101874 [6] Gu Z W, Cheng J, Fu H Z, Zhou K, Hao H Y, Zhao Y T, et al. CE-Net: Context encoder network for 2D medical image segmentation. IEEE Transactions on Medical Imaging, 2019, 38(10): 2281−2292 doi: 10.1109/TMI.2019.2903562 [7] Zhang N, Yu L, Zhang D Z, Wu W D, Tian S W, Kang X J, et al. CT-Net: Asymmetric compound branch transformer for medical image segmentation. Neural Networks, 2024, 170: 298−311 doi: 10.1016/j.neunet.2023.11.034 [8] Lei Y J, Wu Z J, Li Z Y, Yang Y E, Liang Z M. BP-CapsNet: An image-based deep learning method for medical diagnosis. Applied Soft Computing, 2023, 146: Article No. 110683 doi: 10.1016/j.asoc.2023.110683 [9] Patro K K, Allam J P, Neelapu B C, Tadeusiewicz R, Acharya U R, Hammad M, et al. Application of Kronecker convolutions in deep learning technique for automated detection of kidney stones with coronal CT images. Information Sciences, 2023, 640: Article No. 119005 doi: 10.1016/j.ins.2023.119005 [10] Lin J, Huang X R, Zhou H Y, Wang Y Q, Zhang Q N. Stimulus-guided adaptive transformer network for retinal blood vessel segmentation in fundus images. Medical Image Analysis, 2023, 89: Article No. 102929 doi: 10.1016/j.media.2023.102929 [11] Shaker A, Maaz M, Rasheed H, Khan S, Yang M H, Khan F S. UNETR++: Delving into efficient and accurate 3D medical image segmentation. IEEE Transactions on Medical Imaging, 2024, 43(9): 3377−3390 doi: 10.1109/TMI.2024.3398728 [12] Wei J H, Zhu G J, Fan Z, Liu J C, Rong Y B, Mo J J, et al. Genetic U-Net: Automatically designed deep networks for retinal vessel segmentation using a genetic algorithm. IEEE Transactions on Medical Imaging, 2022, 41(2): 292−307 doi: 10.1109/TMI.2021.3111679 [13] Fang X, Yan P K. Multi-organ segmentation over partially labeled datasets with multi-scale feature abstraction. IEEE Transactions on Medical Imaging, 2020, 39(11): 3619−3629 doi: 10.1109/TMI.2020.3001036 [14] Zhang T L, Wang X Z. Anchorwise fuzziness modeling in convolution-transformer neural network for left atrium image segmentation. IEEE Transactions on Fuzzy Systems, 2024, 32(2): 398−408 doi: 10.1109/TFUZZ.2023.3298904 [15] Kumar A, Fulham M, Feng D G, Kim J. Co-Learning feature fusion maps from PET-CT images of lung cancer. IEEE Transactions on Medical Imaging, 2020, 39(1): 204−217 doi: 10.1109/TMI.2019.2923601 [16] Zhou Y, Ahmed T S, Wang M, Newman E A, Schmetterer L, Fu H Z, et al. Masked vascular structure segmentation and completion in retinal images. IEEE Transactions on Medical Imaging, 2025, 44(6): 2492−2503 doi: 10.1109/TMI.2025.3538336 [17] Wu Y C, Xia Y, Song Y, Zhang D H, Liu D N, Zhang C Y, et al. Vessel-Net: Retinal vessel segmentation under multi-path supervision. In: Proceedings of the 22nd International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI). Shenzhen, China: Springer, 2019. 264-272 [18] Xiang D H, Zhang B, Lu Y X, Deng S M. Modality-specific segmentation network for lung tumor segmentation in PET-CT images. IEEE Journal of Biomedical and Health Informatics, 2023, 27(3): 1237−1248 doi: 10.1109/JBHI.2022.3186275 [19] Yuan Y C, Zhang L, Wang L T, Huang H Y. Multi-level attention network for retinal vessel segmentation. IEEE Journal of Biomedical and Health Informatics, 2022, 26(1): 312−323 doi: 10.1109/JBHI.2021.3089201 [20] Ma Y X, Wang S, Hua Y, Ma R H, Song T, Xue Z G, et al. perceptual data augmentation for biomedical coronary vessel segmentation. IEEE/ACM Transactions on Computational Biology and Bioinformatics, 2023, 20(4): 2494−2505 doi: 10.1109/TCBB.2022.3188148 [21] Hatamizadeh A, Hosseini H, Patel N, Choi J, Pole C C, Hoeferlin C M, et al. RAVIR: A dataset and methodology for the semantic segmentation and quantitative analysis of retinal arteries and veins in infrared reflectance imaging. IEEE Journal of Biomedical and Health Informatics, 2022, 26(7): 3272−3283 doi: 10.1109/JBHI.2022.3163352 [22] Hu K, Jiang S, Zhang Y, Li X Y, Gao X P. Joint-Seg: Treat foveal avascular zone and retinal vessel segmentation in OCTA Images as a joint task. IEEE Transactions on Instrumentation and Measurement, 2022, 71: Article No. 4007113 doi: 10.1109/tim.2022.3193188 [23] McKetty M H. The AAPM/RSNA physics tutorial for residents. X-ray attenuation. RadioGraphics, 1998, 18(1): 151−163 doi: 10.1148/radiographics.18.1.9460114 [24] Long J, Shelhamer E, Darrell T. Fully convolutional networks for semantic segmentation. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR). Boston, USA: IEEE, 2015. 3431-3440 [25] Ronneberger O, Fischer P, Brox T. U-Net: Convolutional networks for biomedical image segmentation. In: Proceedings of the 18th International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI). Munich, Germany: Springer, 2015. 234-241 [26] Zhou Z W, Rahman Siddiquee M M, Tajbakhsh N, Liang J M. UNet++: A nested U-Net architecture for medical image segmentation. In: Proceedings of the 4th International Workshop on Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support. Granada, Spain: Springer, 2018. 3-11 [27] Alom M Z, Yakopcic C, Taha T M, Asari V K. Nuclei segmentation with recurrent residual convolutional neural networks based U-Net (R2U-Net). In: Proceedings of the IEEE National Aerospace and Electronics Conference (NAECON). Dayton, USA: IEEE, 2018. 228-233 [28] Jha D, Riegler M A, Johansen D, Halvorsen P, Johansen H D. DoubleU-Net: A deep convolutional neural network for medical image segmentation. In: Proceedings of the 33rd International Symposium on Computer-Based Medical Systems (CBMS). Rochester, USA: IEEE, 2020. 558-564 [29] Xiao X, Lian S, Luo Z M, Li S Z. Weighted Res-UNet for high-quality retina vessel segmentation. In: Proceedings of the 9th International Conference on Information Technology in Medicine and Education (ITME). Hangzhou, China: IEEE, 2018. 327-331 [30] Isensee F, Jaeger P F, Kohl S A A, Petersen J, Maier-Hein K H. nnU-Net: A self-configuring method for deep learning-based biomedical image segmentation. Nature Methods, 2021, 18(2): 203−211 doi: 10.1038/s41592-020-01008-z [31] Oktay O, Schlemper J, Folgoc L L, Lee M C H, Heinrich H P, et al. Attention U-Net: Learning where to look for the pancreas. arXiv preprint arXiv: 1804.03999, 2018. [32] Fan D P, Ji G P, Zhou T, Chen G, Fu H Z, Shen J B, et al. PraNet: Parallel reverse attention network for polyp segmentation. In: Proceedings of the 23rd International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI). Lima, Peru: Springer, 2020. 263-273 [33] Tomar N K, Jha D, Riegler M A, Johansen H D, Johansen D, Rittscher J, et al. FANet: A feedback attention network for improved biomedical image segmentation. IEEE Transactions on Neural Networks and Learning Systems, 2023, 34(11): 9375−9388 doi: 10.1109/TNNLS.2022.3159394 [34] Ding W P, Wang H P, Huang J S, Ju H R, Geng Y, Lin C T, et al. FTransCNN: Fusing transformer and a CNN based on fuzzy logic for uncertain medical image segmentation. Information Fusion, 2023, 99: Article No. 101880 doi: 10.1016/j.inffus.2023.101880 [35] Wang H N, Cao P, Wang J Q, Zaiane O R. UCTransNet: Rethinking the skip connections in U-Net from a channel-wise perspective with transformer. In: Proceedings of the 36th AAAI Conference on Artificial Intelligence. Palo Alto, USA: AAAI Press, 2022. 2441-2449 (查阅网上资料, 未找到本条文献出版地信息, 请核对) [36] Cao H, Wang Y Y, Chen J, Jiang D S, Zhang X P, Tian Q, et al. Swin-Unet: Unet-like pure transformer for medical image segmentation. In: Proceedings of the European Conference on Computer Vision. Tel Aviv, Israel: Springer, 2022. 205-218 [37] Hu W, Yang S Y, Guo W F, Xiao N, Yang X P, Ren X Y. STC-UNet: Renal tumor segmentation based on enhanced feature extraction at different network levels. BMC Medical Imaging, 2024, 24(1): Article No. 179 doi: 10.1186/s12880-024-01359-5 [38] Ruan J C, Li J C, Xiang S C. VM-UNet: Vision mamba UNet for medical image segmentation. ACM Transactions on Multimedia Computing, Communications and Applications, to be published, DOI: 10.1145/3767748 [39] Chen T X, Zhou X D, Tan Z T, Wu Y, Wang Z Y, Ye Z, et al. Zig-RiR: Zigzag RWKV-in-RWKV for efficient medical image segmentation. IEEE Transactions on Medical Imaging, 2025, 44(8): 3245−3257 doi: 10.1109/TMI.2025.3561797 [40] Ma J, He Y T, Li F F, Han L, You C Y, Wang B. Segment anything in medical images. Nature Communications, 2024, 15(1): Article No. 654 doi: 10.1038/s41467-024-44824-z [41] Jiao R S, Liu Q P, Zhang Y, Pu B Z, Xue B S, Cheng Y, et al. RECIST.Surv: Hybrid multi-task transformer for hepatocellular carcinoma response and survival evaluation. IEEE Transactions on Image Processing, 2025, 34: 3873−3888 doi: 10.1109/TIP.2025.3579200 [42] Zhao L X, Wang T, Chen Y B, Zhang X L, Tang H, Lin F X, et al. A novel framework for segmentation of small targets in medical images. Scientific Reports, 2025, 15(1): Article No. 9924 doi: 10.1038/s41598-025-94437-9 [43] Ahmadi M, Biswas D, Lin M H, Vrionis F D, Hashemi J, Tang Y F. Physics-informed machine learning for advancing computational medical imaging: Integrating data-driven approaches with fundamental physical principles. Artificial Intelligence Review, 2025, 58(10): Article No. 297 doi: 10.1007/s10462-025-11303-w [44] Wang X L, Girshick R, Gupta A, He K M. Non-local neural networks. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition. Salt Lake City, USA: IEEE, 2018. 7794-7803 [45] Huang Z L, Wang X G, Huang L C, Huang C, Wei Y C, Liu W Y. CCNet: Criss-cross attention for semantic segmentation. In: Proceedings of the IEEE/CVF International Conference on Computer Vision (ICCV). Seoul, Korea: IEEE, 2019. 603-612 [46] Ruan J C, Xiang S C, Xie M Y, Liu T, Fu Y Z. MALUNet: A multi-attention and light-weight UNet for skin lesion segmentation. In: Proceedings of the IEEE International Conference on Bioinformatics and Biomedicine (BIBM). Las Vegas, USA: IEEE, 2022. 1150-1156 [47] Wu Y X, Liao K L, Chen J T, Wang J H, Chen D Z, Gao H H, et al. D-Former: A U-shaped dilated transformer for 3D medical image segmentation. Neural Computing and Applications, 2023, 35(2): 1931−1944 doi: 10.1007/s00521-022-07859-1 [48] Imtiaz T, Fattah S A, Kung S Y. BAWGNet: Boundary aware wavelet guided network for the nuclei segmentation in histopathology images. Computers in Biology and Medicine, 2023, 165: Article No. 107378 doi: 10.1016/j.compbiomed.2023.107378 [49] Bernard O, Lalande A, Zotti C, Cervenansky F, Yang X, Heng P A, et al. Deep learning techniques for automatic MRI cardiac multi-structures segmentation and diagnosis: Is the problem solved?. IEEE Transactions on Medical Imaging, 2018, 37(11): 2514−2525 doi: 10.1109/TMI.2018.2837502 [50] Xu Y S, Tang J Q, Men A D, Chen Q C. EviPrompt: A training-free evidential prompt generation method for adapting segment anything model in medical images. IEEE Transactions on Image Processing, 2024, 33: 6204−6215 doi: 10.1109/TIP.2024.3482175 [51] Zhou S H, Nie D, Adeli E, Yin J P, Lian J, Shen D G. High-resolution encoder-decoder networks for low-contrast medical image segmentation. IEEE Transactions on Image Processing, 2020, 29: 461−475 doi: 10.1109/TIP.2019.2919937 [52] Qiu Z X, Hu Y, Chen X S, Zeng D, Hu Q Y, Liu J. Rethinking dual-stream super-resolution semantic learning in medical image segmentation. IEEE Transactions on Pattern Analysis and Machine Intelligence, 2024, 46(1): 451−464 doi: 10.1109/TPAMI.2023.3322735 [53] Zhang T Y, Zheng S M, Cheng J, Jia X, Bartlett J, Cheng X X, et al. Structure and intensity unbiased translation for 2D medical image segmentation. IEEE Transactions on Pattern Analysis and Machine Intelligence, 2024, 46(12): 10060−10075 doi: 10.1109/TPAMI.2024.3434435 [54] Bilic P, Christ P, Li H B, Vorontsov E, Ben-Cohen A, Kaissis G, et al. The liver tumor segmentation benchmark (LiTS). Medical Image Analysis, 2023, 84: Article No. 102680 doi: 10.1016/j.media.2022.102680 [55] Lyu F, Ye M, Ma A J, Yip T C F, Wong G L H, Yuen P C. Learning from synthetic CT images via test-time training for liver tumor segmentation. IEEE Transactions on Medical Imaging, 2022, 41(9): 2510−2520 doi: 10.1109/TMI.2022.3166230 [56] Heller N, Sathianathen N, Kalapara A, Walczak E, Moore K, Kaluzniak H, et al. The KiTS19 challenge data: 300 kidney tumor cases with clinical context, CT semantic segmentations, and surgical outcomes. arXiv preprint arXiv: 1904.00445, 2019. [57] 廖苗, 杨睿新, 赵于前, 邸拴虎, 杨振. 基于CE-TransNet的腹部CT图像多器官分割. 自动化学报, 2025, 51(6): 1371−1387Liao Miao, Yang Rui-Xin, Zhao Yu-Qian, Di Shuan-Hu, Yang Zhen. Multi-organ segmentation from abdominal CT images based on CE TransNet. Acta Automatica Sinica, 2025, 51(6): 1371−1387 [58] 贾熹滨, 郭雄, 王珞, 杨大为, 杨正汉. 一种迭代边界优化的医学图像小样本分割网络. 自动化学报, 2024, 50(10): 1988−2001Jia Xi-Bin, Guo Xiong, Wang Luo, Yang Da-Wei, Yang Zheng-Han. A few-shot medical image segmentation network with iterative boundary refinement. Acta Automatica Sinica, 2024, 50(10): 1988−2001 [59] Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. In: Proceedings of the 3rd International Conference on Learning Representations (ICLR). San Diego, USA: ICLR, 2015. [60] He K M, Zhang X Y, Ren S Q, Sun J. Deep residual learning for image recognition. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR). Las Vegas, USA: IEEE, 2016. 770-778 [61] Huang G, Liu Z, Van Der Maaten L, Weinberger K Q. Densely connected convolutional networks. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR). Honolulu, USA: IEEE, 2017. 4700-4708 -
计量
- 文章访问数: 137
- HTML全文浏览量: 110
- 被引次数: 0



 下载:
下载: